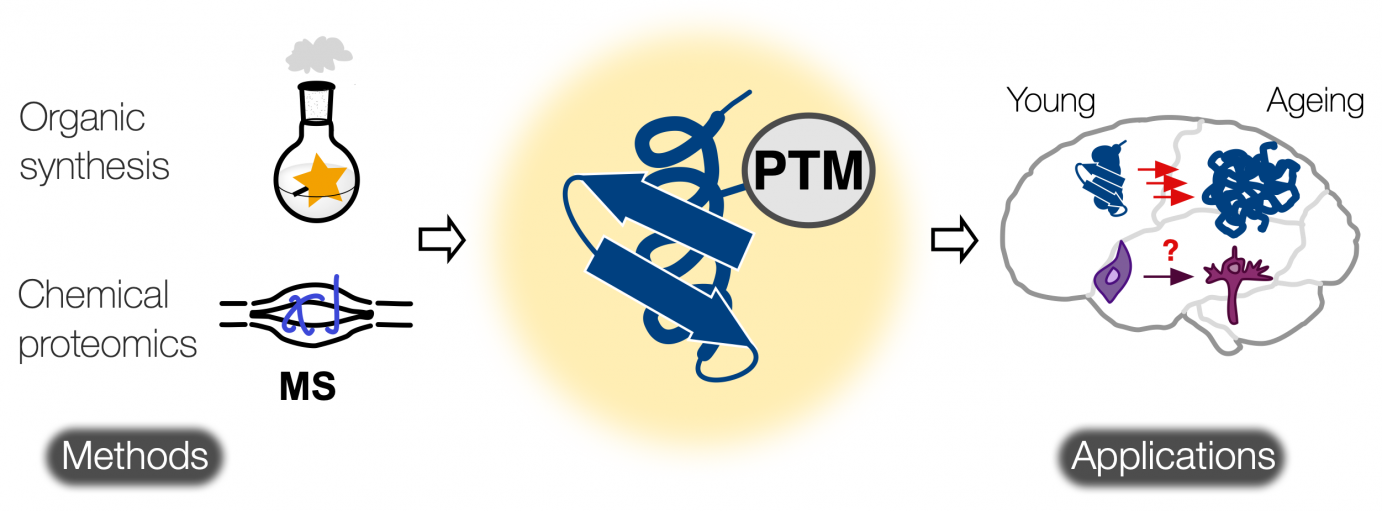
Chemical Biology of Protein Post-translational Modifications
Tweaking protein post-translational modifications to tackle neurological diseases.
Protein post-translational modifications (PTMs) create a complex regulatory layer of proteins’ activity providing the cell with mechanisms to swiftly adjust its metabolism to changing environment, and hence keep the cell homeostasis. Central processes modulating neurodevelopment and neurodegeneration are controlled by protein PTMs. Our objectives are to map and functionally characterize protein PTMs to better understand the fundamental regulatory pathways behind neurodevelopment and to identify PTM-based markers for early diagnosis of neurodegenerative diseases. We aim to develop creative chemical biology tools, chemical proteomic methods and mass spectrometry techniques for characterization of protein PTMs in neurological disorders.
Selected publications:
Hoffmann, L.; Eckl, E.-M.; Bérouti, M.; Pries, M.; Koller, A.; Guhl, C.; Hellmich, U. A.; Hornung, V.; Xiang, W.; Jae, L. T.; Kielkowski, P.: "AMPylation regulates 5'-3' exonuclease PLD3 processing." Mol. Cell. Proteomics, 2025, DOI: 10.1016/j.mcpro.2025.101051.
Chen, H.; Zang, L.; Kielkowski, P.: "Self-assembled PROTACs enable glycoproteins degradation in the living cells." Chem. Sci., 2025, 16, 8060-8068. DOI: 10.1039/D5SC00400D.
Wiest, A.; Kielkowski, P.: "Cu-Catalyzed Azide−Alkyne−Thiol Reaction Forms Ubiquitous Background in Chemical Proteomic Studies." JACS, 2024, 3, 2151-2159. DOI: 10.1021/jacs.3c11780.
Makarov, D.; Kielkowski, P.: "Chemical proteomics reveals protein tyrosination beyond the alpha-tubulins in human cells." ChemBioChem, 2022, DOI: 10.1002/cbic.202200414.
Becker, T.; Wiest, A.; Telek, A.; Bejko, D.; Hoffman-Röder, A.; Kielkowski, P.: "Transforming chemical proteomics enrichment into high throughput method using SP2E workflow." JACS Au, 2022 DOI: 10.1021/jacsau.2c00284.
Becker, T.; Cappel, C.; Di Matteo, F.; Sonsalla, G.; Kaminska, E.; Spada, F.; Cappello, S.; Damme, M.; Kielkowski, P.: "AMPylation profiling during neuronal differentiation reveals extensive variation on lysosomal proteins." iScience, 2021, 24, 103521. DOI: 10.1016/j.isci.2021.103521.
Sieber, S. A.; Cappello, S.; Kielkowski, P.: "From young to old: AMPylation hits the brain." Cell Chem. Biol., 2020, 27, 773-779 (The Perspective highlighted on the Cover page).
Kielkowski, P.; Buchsbaum, I.; Kirsch, V.; Bach, N.; Drukker, M.; Cappello, S.; Sieber, S.: "FICD activity and AMPylation remodelling modulate human neurogenesis." Nat. Commun., 2020, 11, 517.